|
Using emulsion PCR amplificiation, it creates many copies of a DNA strand attached to a bead. Available in the mid 2000s, is similar to Sanger Sequencing but is done in parallel and requires far less preparation. They hope to read an entire genome in under an hour for $100.Ĥ54 (700bp) by Roche uses pyrosequencing. This fluorscence is recorded and a DNA sequence is discerned. It uses polymerase to add phospholinked nucleotides which carry fluorescent labels on the terminal phosphate, which is cleaved away during replication. PacBio (2-10kb) uses single molecule real time sequencing. Illumina is currently the most widely-employed technology in DNA sequencing. This is done in parallel, with 150 million DNA fragments per flow cell. 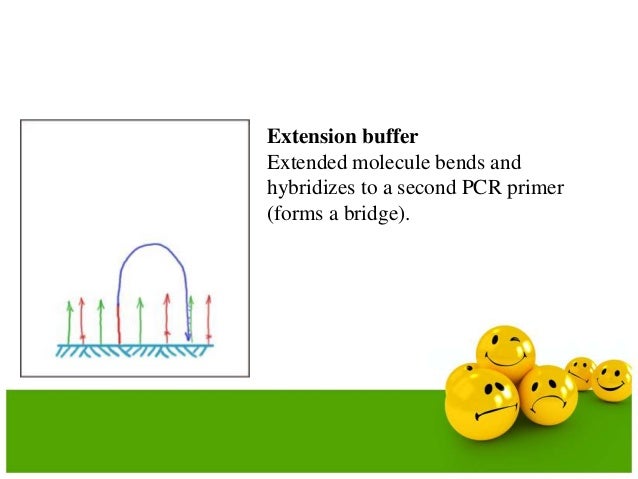
As each base is added a camera records the fluorescent signal emitted to determine the sequence. 
Bridge amplification is employed to sythesize copies of the DNA on the surface of a flow cell. A certain size fragment is selected and amplified through PCR. 
DNA which has been sheared into small fragments is separated on gel by fragment size. It is another method of chain termination. Illumina (150bp) performs sequencing by synthesizing complementary DNA strands. However, this method of sequencing requires a lot of work and large amount of reagent, making the bases sequenced per dollar expensive - nearly 50,000 times more expensive per base than Illumina sequencing.įor more information on chain termination sequencing, there are YouTube videos which illustrate it in more depth, as well as good information at JGI.Īlso see the 3-part series of shotgun sequencing: part I, part II and part III. The ddNTPs will be added in random places along each strand, and by stopping the polymerase action randomly we can run the various DNA fragments out on gels and visualize the chain-terminations to determine the DNA sequence.Ĭhain termination sequencing is useful as it can read long sequences of DNA (over 800bp) and is very accurate. This allows us to to see only one color to infer the nucleotide. The dNTPs and ddNTPs are added into four separate containers with a radioactive primer. The polyermase enzyme will recognize the ddNTP is not a correct dNTP, so will cease adding nucleotides at that location. In chain termination we use both dNTPs and ddNTPs (dideoxyribonucleotide triphosphates, or free A, C, G and T nucleotides, where the hydroxyl group on the third carbon is missing an oxygen) together. Sanger Sequencing, or capillary sequencing, is a way of reading the nucleotides by using chain termination.ĭNTPs are the free nucleotides (deoxyribonucleotide triphosphates, or free A, C, G and T nucleotides) dATP, dCTP, dGTP and dTTP. We use an ABI 3730xl, a type of Sanger sequencer. TempliPhi (an enzyme, free nucleotides and hexamers) is added to the wells. The plasmids can then be amplified using rolling circle amplification. The TE buffer will protect the DNA while we heat up the plate to 95C, allowing us to separate the plasmid from the bacteria by bursting the cell walls. 
We can then use a robotic colony picker to create new plates for the white colonies, as white colonies represent bacteria containing our fragments.Īfter letting the bacteria multiply for a day, we transfer the bacteria into a new 384-well plate and fill each well with a TE buffer. By using an agar plate with X-gal and ampicillin, we can see which colonies have inserted DNA (intact lacZ genes will express a blue color, indicating they obtained a pUC18 plasmid but did not insert our DNA fragment) and the ampicillin will kill any bacteria which did not insert a plasmid. Bacteria which take in the DNA are then referred to as "transformed." Only one in 1000 E. It uses an electric shot to create tiny holes in the bacteria for the plasmids to enter. To induce plasmids to insert into bacteria faster, we can use an electroporlator. The blunt-ended DNA fragments can be inserted into the LacZ gene, disrupting the blue color. By using an engineered plasmid called pUC18, we can use two important traits: the LacZ gene will express a blue color in the presence of X-gal (a sugar analog of lactose), and an ampicillin-resistant gene. This is what is used in cloning.īacteria are great for sorting and copying our DNA. Bacteria routinely pass fragments of DNA to each other through conjugation, in a process known as horizontal gene transfer. We can use bacteria to store DNA and copy DNA.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
